|
For The Agency | For The Technology Developer | For The Magazine Publisher | For The Individual |
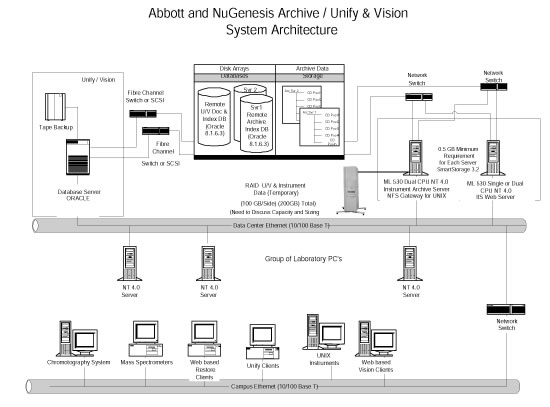
High Performance Computing in the Life SciencesAutomated instrumentation is pushing biochemical science into the supercomputer world. The following is a manuscript for an article published in R&D magazine. R&D magazine holds the copyright for the finished article. C.G. Masi, Contributing Editor "With the advent of high-throughput automated molecular-biology techniques," says Dan Stevens, Market Manager for Life Sciences and Chemistry Computing at supercomputer manufacturer SGI in Yardley, Pennsylvania, "the ability to generate large amounts of biochemical data have become realized in the drug discovery arena. Now, the objective is to turn these data into knowledge and, ultimately, products." Supercomputers can help researchers in three fundamental ways: through bioinformatics, chemical analysis, and chemical modeling. Bioinformatics generally covers the use of database analysis to organize and interpret large collections of research data. These data may come from chemical analysis, modeling, clinical trials, and even public records, such as reports in scholarly journals. Chemical analysis includes all of the techniques needed to interpret data collected by automated laboratory systems. These data come from a variety of analytical techniques, such as mass spectrometry, nuclear magnetic resonance (NMR) and high-performance liquid chromatography (HPLC). Its output is the chemical makeup of samples, which can be used to identify active compounds that might make promising new drugs. Chemical modeling uses a priori physico-chemical modeling techniques to help understand how the active compounds fit together (structural analysis) and interact (dynamics). The supercomputers used are made up of computer hardware, software, data storage and networking elements as shown in Fig. A. What sets them apart from more mundane installations is the size (measured by the number of redundant elements, such as processors) and speed (measured by the number of operations per second) of the aggregate machine, and the high-speed interconnections needed to keep all the components working together at their peak effectiveness.
The main software differences between supercomputers and other computers appears in the application programs they run. Operating systems used are generally variants on UNIX, which has been run on machines of all sizes. The sizes of the problems these computers are generally called upon to address make writing specialized application programs both desirable and resource effective. For example, while almost any modern computer can use spring-and-ball models to simulate the dynamics of small molecules consisting of a few atoms, only the most powerful machines can do the same with typical biological macromolecules. On the other hand, to comprehend the results of the simulation takes a more sophisticated user interface as well. The same can be said of the software needed to set up the problem. High-Performance Applications "There are several different types of customers," says Jeff Augen, Director of Strategy for the IBM's Life Sciences Group in Somers, New York. "Some customers are doing computer-intensive projects that are floating point intensive. They're doing molecular modeling, looking at protein structure, and doing X-ray crystallography. "There are also customers who are doing bioinformatics. They're doing pattern discovery, and searching genomes for matching sequences. That's integer intensive, not floating point intensive, and requires a different type of compute infrastructure. "Finally, there are customers who are building databases of content that they sell or distribute. Content distribution has different requirements for computer infrastructure. It's more like large scale Web serving." Researchers modeling complex molecules need high floating-point performance. They usually work with large parallel supercomputers. An example of that type of machine is IBM's SP-series. Another example would be a cluster of Intel-Pentium-based machines running Linux. Linux is popular because it is a real-time operating system (like UNIX, but unlike Windows) that can easily do embarrassingly parallel problems in a cluster environment. An embarrassingly parallel application is one where the problem itself can be broken down into many, many, many small pieces that can be independently calculated without knowing the results of the calculations of the other pieces. If you take a list of a ten numbers and multiply each one by three, that's a parallel problem. Each calculation is extremely simple and you don't have to know the results of any of the other calculations to get its result. If you have 10 million numbers to multiply by three, that's embarrassingly parallel. It would take forever to do on a single processor, but you could run it on 10 million processors at the same time and get the result very quickly. Then, you just dump all the results into a big database. If you have a thousand different gene sequences and want to search a large database for a match for each one, that, too, is embarrassingly parallel. You can take each of the thousand sequences and, using a thousand processors and one copy of the database in a shared memory, simultaneously search that database for all matches. "You need three basic elements," says Lionel Binns, Worldwide Life and Material Sciences Manager in the High Performance Technical Computing Group at Compaq Computer Corporation in Manchester, U.K. "You need some processing units, you need some interconnect, and you need a huge amount of storage. Those are the three base components of any system that's going to be required to do any protein structural work." "If you're looking at supercomputers capable of doing any work around protein structure, what you're going to end up buying is a very tightly coupled cluster of Alpha-based systems." (See Fig. D.)
The Alpha chip is Compaq's high-performance reduced-instruction-set-computing (RISC) processor. It was originally developed by Digital Equipment Corporation (DEC) in Maynard, Mass. in the 1980s and became part of Compaq's product line when they acquired DEC. "We build a series of machines called the SC series. SC Series computers consist of multiple ES 40 computers, each built around four Alpha processors. These units are then tightly coupled through a very, very fast switch with high bandwidth. Include some special software for it, and you get a very tightly coupled cluster which acts like one system." "In the chemistry market," SGI's Stevens points out, "many of these applications really require large scalable shared memory systems." SGI's Origin 3000 is a non-uniform memory access (NUMA) system providing a large shared memory with extremely high speed I/O. Scalability through modular design is the key to the Origin class servers. Small biotech companies can start with 16 processors, and grow into 128 processor systems. The structure and the infrastructure to support both of those machines is the same because each of the pieces, the processors, the graphics, disk space, I/O, and connectivity, is resident on modules. Historically, people thought of the company's Onyx-class machines as the visualization engines, but you can actually take an Origin class server, put a "G" (which stands for graphics) module in it, and it turns into an Onyx! Integer-Intensive Computing Integer-intensive computing problems in life sciences usually involve holding large DNA or protein sequences in memory, so the machines need a large amount of addressable memory. Usually the problem involves searching those sequences to, for example, match them with some target. Users want to search quickly and repetitively. They may also be sharing the same database across many CPU's, as well. What you need for this type of problem is a storage area network (SAN), where there's a large storage system at the back-end to house the database. Many large CPU's reach out to that storage system to search that same database. They load large pieces of it into their local addressable memory to actually do the searching. Typically, those processors are integer-intensive machines. That is, they are optimized for high performance on integer-arithmetic operations. Their distinguishing feature is high clock speed. The key factor determining the performance of a system like that is how many ways you can split up the problem and how fast the clock runs on those machines. IBM's SP is not the right machine. A better choice would be the company's R6000. These machines run IBM's AIX variant of UNIX. Another good choice might be a cluster of Intel-based machines, if you don't need very large local memory for the sequences. They have high clock speed for fast integer calculations. Integer-intensive calculations also call for fast network performance between the storage device and the CPU's. The processors reach out to the database quite often, unlike those doing floating-point-intensive problems, such as molecular modeling. With so much activity, poor network performance would strongly affect overall machine performance. SANs for these machines usually use high-speed fiber-optic links. The data servers making up the SAN are intelligent storage devices. Their operating systems allows them to serve up data to a large number of CPUs. File-transfer-intensive computing (shown in Fig. C), while it also works with databases, is fundamentally different from integer-intensive operations.
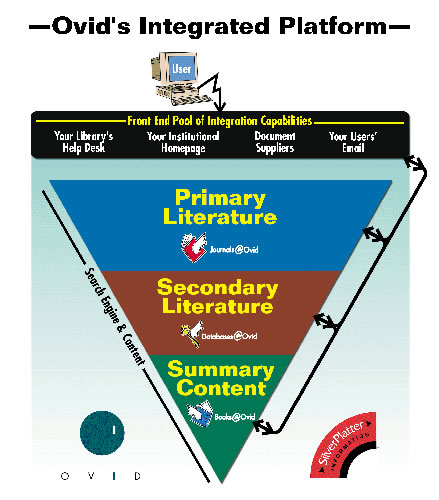
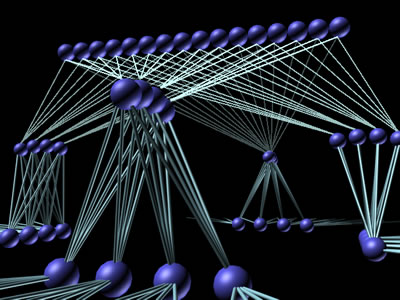
"We have customers," IBM's Augen reports, "who have large databases that they sell access to. Their clients log into their databases and retrieve information." They're combining data from the public databases, from private databases (such as the Celera Human Genome Assembly from Celera Genomics Group, Rockville, Md.), and from databases inside some of their client companies. Their value added includes organizing those data sets so that they can be searched more efficiently. Providing that service requires a lot of precomputing of data on the back end, and then posting it on a server farm that's available across the Internet. The key there is to have the right devices in order to do load balancing between the machine that serves up the data across the Internet. An example of load imbalance is having a thousand customers logged into one server and one customer logged into another, while both servers provide access to the same database. The system has to monitor the load on each individual server and, when one server is becoming overloaded, log the next customer in onto a different machine. At the other end of the system's structure, there is a host of application servers and large storage devices. These take new data being incorporated into the archive and precomputed results of anticipated searches. They post those precomputed results (called metadata) onto the servers on the front end to speed up the process. When a researcher actually requests something that was anticipated, the result will already be there. Massive Databases "If you're a scientist and you need to know something," says Cheryl Chisnell, Vice President for Software Development at Ovid Technologies in New York, N.Y., "you need to go to the literature to get that information. We bring that literature to you, the scientist." Ovid provides a window into the research literature through bibliographic databases, which are indexes to the literature and providing online access to the full text across multiple publishers. Their supercomputer system software is geared towards providing researchers direct access and the ability to query the literature directly. An advantage to using Ovid's library is that the company understands the content of the articles it carries. This understanding, built into the software used to develop metadata for the archive, makes it possible to provide targeted searches that thoroughly cover the information needed without returning a lot of false-positive links to irrelevant articles. They have relationships with publishers of (at present) about 450 scientific journals that allow them to obtain that content, load it into their system, put a search interface and a search engine on top of it, and provide online access to it on a subscription basis. In addition, they will load additional private content for corporate clients who want to make it easier for their own scientists to find their way through their proprietary information. Institutions and companies subscribe to Ovid's databases, then make it available to their internal registered users. So, pharmaceuticals company X designates specific worldwide offices to gain access to the databases. Company X either receives a set of user ID's and passwords to pass out, or else they use IP validation to make sure that their people are the only ones who are getting the access they have purchased. They will decide, of the 450 available journals, to purchase access to, say, 100 of them. Their registered users can then log in from anywhere throughout the world to search those 100 publications. Ovid's databases currently take up about half a terabyte of storage. That includes full text, bibliographic data, metadata, images, etc. This data is stored in a SAN operating in a Sun Solaris environment. There is a pool of front-end machines to serve users calling in, for example via the Internet, and a pool of back end machines. The back end machines are the search engines by interfacing with the repository of information in the SAN. The software to run this operation is also part of Ovid's proprietary technology. They have developed thin-client interfaces allowing registered users to log onto the system with only a Web browser running on their local system. Other proprietary programs search the bibliographic and metadata records for links to relevant articles cataloged into the full-text store. Still other programs are needed to develop the bibliographies and metadata that the search programs will look through in order to find the links that match the users' queries. "Millions and millions of records are created by scientists every year in a typical pharmaceutical company," Gregory Murphy, Vice-President of Enterprise Solutions, NuGenesis Technologies, Westboro, Mass. points out. "What they're trying to do is store them in an organized way in a centralized location. They need to store machine-readable data as well as human-readable data in an organized manner so that it is easily retrievable and usable." NuGenesis builds complete systems to help these companies. They use their proprietary software and various hardware systems. Their software runs on Compaq systems, Windows-NT-based systems or Unix servers. The information is recorded in Oracle databases physically housed in SANs on the back end. "We have roughly 1,200 scientists at a major pharmaceutical account who are storing literally everything that they process in our database," Murphy reports. This particular installation runs on a set of Sun Solaris computers using Oracle databases that are connected to the scientific laboratory's network along with all the analytical instruments. The data that the analytical instruments generate is collected from the laboratory network by a program named "Archive." Archive stores the data actually in a large SAN capable of storing millions of files. The analytical instruments store the data they generate locally on their own disk drives. Archive sweeps those local disk drives for new files, and automatically extracts metadata that describes the raw data. It stores the metadata in an Oracle catalogue, allowing users to find that data at a later date. That metadata describes up to 75 different parameters about each file. Once the system finds the records relating to a query, it can upload the original raw data file. Terascale Computing "We recently sold a very large system to the Pittsburgh Supercomputing Center (PSC) at Carnegie Mellon University in Pittsburgh, Penn.," Compaq's Binns reports. "The completed system will have some 734 nodes, and each node will have four Alpha processors for a total of nearly 3,000 processors." The processors are linked together by a set of very fast high-bandwidth switches called a "fat-tree" switches (see Fig. E), the term "fat tree" indicating that each switch has a high fanout. The switches combine into a matrix called a "federated switch." Multiple fat- tree switches in the matrix are switched together by other switches so that each processor is capable of talking to every other processor within the switched cluster To all intents and purposes, the system will behave like one giant computer.
Although there are hundreds of nodes comprising thousands of processors, each individual node is a consumer-off-the-shelf (COTS) unit. This COTS strategy allows system integrators to buy four-processor computers as individual units, and build them up into very big systems. It is a very cost effective approach to building supercomputers. There must be, of course, a large amount of data storage as well. Each processor will have some storage of its own locally, but there will also be a large SAN serving the entire machine. TCSini, as the system is called, became operational in late December of 2000, well ahead of the February 1 scheduled date, and has seen steadily increasing usage during the January-February-March "friendly user" period, which served as the final round of testing and development. During this period, PSC staff modified scheduling software to accommodate large-scale projects that use all 256 processors. A number of such projects ran during the second half of March, for scheduled periods of dedicated use as long as 12 hours, demonstrating TCSini's ability to support sustained whole-system applications. The PSC modified scheduler makes it possible to integrate such projects into the system workload while still allowing wide access to smaller-scale projects. TCSini will later this year be replaced by the full-scale TCS, a 3,000-processor, six teraflop system that will be the most powerful system in the world available for public research. Blue Gene is an IBM Research project to build a very large supercomputer, ten to a hundred times bigger than what's currently available, and use it to advance the state of the art of simulating biomolecular systems. Announced in December of 1999, the project is expected to be a roughly five-year effort. One class of problems that Blue Gene might attack is to understand the dynamics of protein folding. Other classes of problems might be simulating how small molecules bind to proteins, how substrates bind to enzymes, or how proteins interact with each other. "The idea is," says Dr. Joe Jasinski, Senior Manager of Computational Biology Center at IBM Research in Yorktown Heights, New York, "that the technology for many of these kinds of simulations for small molecule systems fairly well worked out. Two things hold us back from applying them to large systems: computer power, and the accuracy of the models." For Blue Gene to succeed, the researchers need to develop the technology to build computer systems with much larger computational capability than is currently available, and to advance the state of the art of the models. For example, the largest supercomputer in the world today has 12 teraflops peak performance. Blue Gene's goal is to build a machine in the hundred-to-thousand teraflop class in the next four years. That would be ten to one hundred times more computing power than is currently available. The machine will be built in a fundamentally different way from supercomputers are built today. The idea is to use a cellular architecture. Supercomputers are organized today in the same way one normally builds computers. There is a central processing unit, some memory chips, and some communication (I/O) chips on separate boards. Those boards all go into a box along with a disk drive, and that's a computer. The idea behind the cellular architecture is to put all of that functionality (except for the disk drive) onto a single silicon chip. The logic, memory and I/O capability will be integrated on the same chip. That chip becomes a cell. Then you build up the systems from those cells. "We think it has advantages in price/performance, in physical size, and in power consumption," Jasinski says. "One of the things that Blue Gene is trying to test is whether you can build practical systems using that kind of architecture." JA: "If you built a cluster of a million Pentium/Windows-based machines," Augen points out, "you could do the same thing--except for the fact that it would use 14 megawatts of electricity and one of those PCs would crash every tenth of a second. "You would also have to buy a million licenses for Windows from Microsoft. It would take up five acres of physical space and use as much electricity as a small city. "That's what it would be like to build Blue Gene out of the individual PC's running Windows. When you think about building large scale clusters, it doesn't make sense to do it by linking individual PC's together. That's why we have supercomputers like the SP and that's why we're developing Blue Gene. Fig. B: The terascale computing system at the Pittsburgh Supercomputing Center will, when all of its nodes come on line, be the largest private computer system in the world. Courtesy Carnegie Mellon University, Pittsburgh, Penn. For More Information Compaq Computer Corporation IBM Corporation NuGenesis Technologies Ovid Technologies SGI |
|
Home | About Us | Technology Journalism | Technology Trends Library | Online Resources | Contact Us For The Agency | For The Technology Developer | For The Magazine Publisher | For The Individual © , C. G. Masi Technology Communications, Privacy Policy P.O. Box 10640, 978 S. San Pedro Road, Golden Valley, AZ 86413, USA Phone: +1 928.565.4514, Fax: +1 928.565.4533, Email: cgmasi@cgmasi.com, Web: www.cgmasi.com Developed by Telesian Technology Inc. |